Problem 32: Some DNA can jump.
Explore other organisms with transposable elements to determine how to locate genes using a transposon.
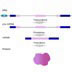
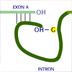
HI! Corn isn't the only organism with transposable elements. In a method called tranposon-tagging, scientists use transposons as location markers to find genes they're interested in. Suppose a scientist is interested in wing formation in flies. She can use transposons to locate genes involved in wing formation by introducing a transposon and looking for wing mutants. For the transposon to affect the organism's phenotype, where in the gene must it insert? Anywhere within a gene. No, a transposon can insert anywhere but won't always have an effect. Anywhere within an intron. No, inserting within an intron won't have an effect, unless the insertion disrupts the exon/intron splice junctions. Both of the above. No, inserting within an intron won't have an effect, unless the insertion disrupts the exon/intron splice junctions. Anywhere within an exon or the control region of the gene. That is correct. In order for the transposon to alter phenotype, it has to jump into an exon or the control region of the gene. Inserting into a non-coding region like an intron will have no effect, unless the insertion disrupts the exon/intron splice junctions. Even inserting into an exon may not have a noticeable effect. It all depends on how dissimilar the mutant protein is from the original. Most organisms have inactive transposons; they're missing the transposase enzyme, so the element no longer jumps. However, the transposase gene can be crossed into a stock, in which case the F1 progeny will have transposons in new positions. If you examine the F1 population of such a cross, what can you expect? All the F1 progeny will have mutations caused by transposon insertions. No, not all transposon insertions cause mutations. All the mutations caused by transposon insertions will have visible phenotypes. No, since this is F1, only insertions that cause dominant mutations will have visible phenotypes. There will be some F1 progeny with visible phenotypes due to transposon insertions. That is correct. None of the above; there are no mutations and no visible phenotypes until F2. No, there will be transposon insertions in F1 that cause visible phenotypes. The F1 population will contain organisms with mutations caused by transposon insertions. However, you will only see a visible phenotype if the mutation is dominant. To see a recessive phenotype, a tranposon would have to insert into the same place on both chromosomes, and this is unlikely. You pick an interesting mutant from the F1 progeny and you want to find the gene responsible for the mutation. Remembering that this is a dominant mutation, what do you have to do next? Make sure the mutation breeds true. Yes, you need to make sure that the transposon doesn't jump out again. Cross the mutant to a "wild-type" strain to get rid of other transposon-induced mutations. Yes, you need to cross out other transposon-induced mutations that have nothing to do with your gene. Both of the above. That is correct. None of the above, the gene is transposon-tagged. Yes, but there are other transposon copies that must be dealt with. Because the transposase gene is present, a transposon can continue to jump. Also, there are many other transposon copies within your mutant. Crossing the mutant with a "wild-type" strain — one without the transposase and thus less transposon copies — and then backcrossing can address both these problems. You backcross the mutant several times. You isolate DNA from both the mutant and the wild-type. You cut the DNA with a restriction enzyme and run the samples on a gel. Assuming that you were successful in removing all the other transposons, what do you see on the gel? Remember this is genomic DNA. Both lanes will have distinct bands. The mutant lane will have an "extra" band. No, you won't see distinct bands; genomic DNA cut with one enzyme gives too many fragments. Both lanes will have a smear; there won't be any distinct band. That is correct. Both lanes will have distinct bands and look exactly the same. No, you won't see distinct bands; genomic DNA cut with one enzyme gives too many fragments. Cutting genomic DNA with one restriction enzyme will give many different-sized fragments. When run out on a gel, you will see a smear pattern without any distinctive bands. The DNA fragment with the transposon-tagged gene is present in the mutant sample. A technique called Southern blotting can be used to identify the fragment. In Southern blotting, the DNA on a gel is transferred and fixed onto a nylon membrane. A radioactive DNA fragment, called a probe, hybridizes to specific fragments on the membrane, thus identifying the fragment of interest. Most organisms have a number of "permanent" transposon copies in their genome that do not jump. Using transposon DNA as the radioactive probe, a Southern blot will identify all the permanent transposons in the wild-type and the mutant, along with any new copies in the mutant. Courtesy of Dr. R. Steven, University of Toronto. Noncommercial, educational use only. This is a Southern blot from an actual transposon-tagging experiment. The organism is Caenorhabditis elegans, the transposon is Tc1, and the tagged gene is unc-73. Wild-type C. elegans has more than 20 permanent Tc1 copies. The first lane is wild-type DNA, the next eight lanes have DNA from transposon-tagged strains of unc-73. The last two lanes have DNA from a different mutation, dpy-5, not caused by a transposon. Click on the arrow that points to the position of the Tc1-tagged unc-73 gene. Arrow 1: No, all the lanes have this band. This is one of the permanent Tc1 copies. Arrow 2: No, there seems to be an extra band in lanes 6-8, but it's not in all the Tc1-tagged unc-73 lanes. Arrow 3: Correct Arrow 4: No, all the lanes have this band. This is one of the permanent Tc1 copies. This extra band is present in all the Tc1-tagged unc-73 mutants and absent in the wild-type or dpy-5 mutants. This band must be the Tc1 that jumped into and caused a mutation in the unc-73 gene. CONGRATULATIONS!!! YOU'RE SO SMART!
transposon insertions, intron splice, splice junctions, transposable elements, exon, control region, transposase, Southern blot, C elegans
- ID: 16686
- Source: DNALC.DNAFTB
Related Content
15549. Transcription/translation - Exons and introns
In most eukaryotic genes, coding regions (exons) are interrupted by noncoding regions (introns).
16938. 3D Animation of RNA Splicing
An animation of the crucial RNA editing step called splicing.
16586. Problem 26: RNA was the first genetic molecule.
Explore how RNA can self-splice.
16949. Antisense Oligonucletotide Therapy for SMA
Dr. Krainer explains the science behind antisense therapy for SMA.
16941. 2D Animation of Alternative RNA Splicing
An animation shows alternate splicing of the SMN2 gene.
16528. Concept 24: The RNA message is sometimes edited.
RNA splicing removes non-coding introns and splices together exons.
16937. An Explanation of RNA Splicing
Dr. Sharp explains the process of RNA splicing.
16948. Alternative RNA Splicing Therapy for Spinal Muscular Atrophy
Drs. Sharp and Sumner describe how RNA splicing can be used as a therapy for SMA.
16933. 3D Animation of DNA to RNA to Protein
An animation shows how the DNA genetic "code" is made into protein.
16939. RNA Splicing
A step-by-step 2D animation shows the details of RNA splicing.