Gallery 36: Differential Expression Data
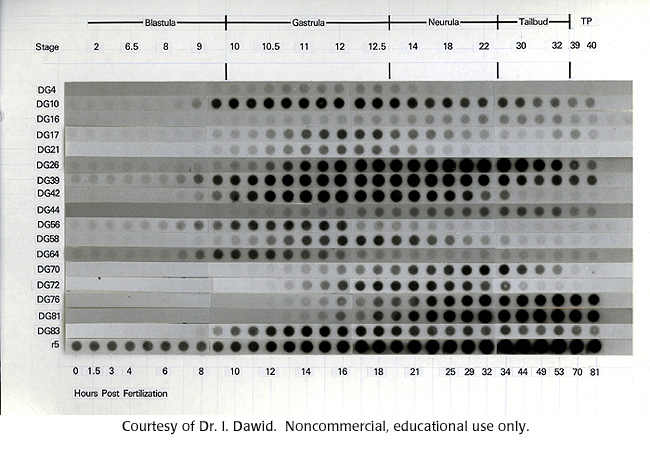
Figure from Sargent and Dawid's differential expression experiment in frog embryos.
differential expression, expression data, dawid, sargent, differential, frog
- ID: 16740
- Source: DNALC.DNAFTB
Related Content
16754. Biography 36: Igor Dawid (1935- )
Igor Dawid did some of the first differential gene expression studies using cDNA subtraction.
16755. Biography 36: Thomas Dean Sargent (1953- )
Tom Sargent did some of the first differential gene expression studies using cDNA subtraction.
16736. Animation 36: Different genes are active in different kinds of cells.
Igor Dawid and Thomas Sargent explain how they developed subtractive mRNA hybrization to find genes expressed by different cell types. Pat Brown and Steve Fodor show how genomes can be screened with DNA arrays and GeneChips™
16743. Video 36: Tom Sargent, clip 1
Part I: Theories on how organisms end up with differentiated cells.
16744. Video 36: Tom Sargent, clip 2
Part II: Theories on how organisms end up with differentiated cells.
16746. Video 36: Tom Sargent, clip 4
Sargent's gene library and comparisons with today's gene chips.
16747. Video 36: Tom Sargent, clip 5
Molecules that regulate development are similar in different organisms.
16748. Video 36: Tom Sargent, clip 6
Why use other animal models to study development?
10923. "Outcome of differential fecundity"
"Outcome of differential fecundity"
16745. Video 36: Tom Sargent, clip 3
The cell as a micro-computer that can interpret signals for different responses.