Gene knockout in mice
This method uses homologous recombination to disable a gene of interest to produce a genetic knockout.
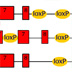
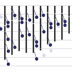
This brown or agouti colored mouse is the type of mouse that we derive embryonic stem cells from. When this mouse becomes pregnant, the embryos are collected, and within those embryos are a number of cells called the inner cell mass. 237 (Pull out several embryos. These can be simply extracted from the embryo and placed into a tissue culture condition where those cells are fed a medium, and they grow on a plastic surface. This is very similar to growing bacteria on a bacterial plate. 224 (Move pipet into embryo and continue to culture dish. Let me tell you how we create a mutation in a gene in a mouse. This gene is PSD-95, also known as post-synaptic density 95, which we later found to be important for learning and memory. 196 (Remove culture dish from this series. We take a piece of DNA from this gene, and we modify it by inserting a neo, or neomycin, cassette into it. This disrupts the gene so that it could not ultimately express normal protein. This vector we then make a second modification to – which is to add a diphtheria toxin cassette to the end of the vector. This DNA vector we introduce into the embryonic stem cells. The way we do this is to take the embryonic stem cells and simply mix them with the DNA, then give them an electric shock. That causes the DNA to be taken into some of those cells. This process is called electroporation. (Add in culture dish below. Pull out cell from culture dish, as in next scene, and show a vector going in a disappearing.) After electroporation, there are three types of events that occur. The first is homologous recombination. That is when the DNA vector finds the correct gene inside the embryonic stem cells, and integrates into that gene and replaces it. When it does so, it loses the diphtheria toxin cassette that was on the end of the vector, but retains the neomycin cassette. In this situation the vector has homologously recombined with the endogenous gene. The second type of event is when the vector goes into an embryonic stem cell, and it does not find the correct gene, but lands at random and integrates into any part of the chromosome. In this situation the vector integrates and retains the diphtheria toxin gene, as well as the neomycin cassette. The third type of event is when the cells don’t take up any DNA at all, so the chromosomes are left completely unmodified. We now need to sort out the three types of events that have occurred. In the case of the cells that have taken up DNA and randomly integrated it – they have kept the diphtheria toxin gene, which is highly toxic and simply kills those cells. We now need to sort out those cells that have not taken up any DNA at all from those that taken up DNA through homologous recombination. The homologously recombined vector carries the neomycin gene, which gives resistance to an antibiotic. So, we simply put an antibiotic onto the cultures, and that kills off all of those cells that didn’t take up the DNA. We are now left with a plate of growing cells where there are small colonies or small lumps of cells. Each one of the colonies contains cells carrying the correct genetic modification that has occurred through homologous recombination. The embryonic stem cells growing on the plate can be collected and then injected into the blastocyst of a donor embryo. This donor embryo will come from a white mouse, and you will recall that the embryonic stem cells came from a brown mouse. So this embryo, when now put into a surrogate mother, will ultimately give birth to a live pup – which will grow up and have cells from the donor, the white mouse, and cells from the embryonic stem cells, the brown mouse. This animal is called a chimera. The characteristic of a chimera is that there are cells from two types of mice, and you can see that in their coat color. The brown patches of fur come from cells from the embryonic stem cells, and the white patches of fur come from the donor mice. Because the chimera is made up of cells from two kinds of mice, that means they must be made up of chromosomes from two types of mice. There are those chromosomes from the albino donor, and there are those chromosomes that come from the brown embryonic stem cells. The chimera contains cells that are both normal cells from the donor and embryonic stem cells that carry the mutation. We want mice that carry only the mutation. To get those mice, we take the chimera and we breed it with some normal mice. In the progeny of that cross we will find some mice that carry the mutation on one chromosome. These are mice that have shown germline transmission and carry a heterozygous mutation in the PSD-95 gene. Of the brown mice that are born from the chimera breeding, we need to sort out those that carry the gene mutation that we engineered from those that do not. For this we use a simple kind of DNA analysis, where we extract the DNA from a piece of tissue from these mice, and we test it using a PCR, or a polymerase chain reaction, experiment. In this experiment, we choose to use two different primers, and one of those primers is within the neomycin, or neo, cassette. So when we do the PCR reaction, we can only detect a DNA fragment from the genetically modified mice, and we cannot detect one from the mice that are not genetically modified. The PCR reaction used in this example generates a 400 base pair fragment of DNA that is unique to the genetically modified chromosomes. 17B This can be readily visualized using electrophoresis of the DNA fragments. Here are examples from four different mice where there are bands observed in only two of those four lanes. It is those lanes, where the band is observed, that indicate that that DNA came from the genetically modified mice. In the case of PSD-95, and other autosomal genes, the type of genetic modification that I’ve just described results in the deletion or mutation of only one of the two different chromosomes encoding that gene. What we would like to achieve is mice that have a mutation in both copies of those genes. For this purpose we simply breed together two of these heterozygous mice and examine the offspring of that breeding to see if those mice are homozygous – or contain mutations in both copies of the gene. We now need to sort out in that litter of mouse pups – from the breeding of the two heterozygotes – those animals that contain a mutation in both copies of the gene, from mice that are heterozygous containing the mutation in only one copy, from mice that contain no mutation at all. For this purpose, we again use a PCR reaction. In this example, the PCR reaction is using primers that are on either side of the neomycin cassette, and, in the case of the genetically modified chromosome, the PCR reaction will result in a fragment 1200 base pairs long. Those primers on a normal chromosome, or a wild type, chromosome produce a shorter fragment of 400 base pairs long. We now apply that PCR reaction to DNA taken from the mice of a heterozygous intercross. We run that PCR reaction out on an electrophoresis gel and simply look at the pattern of band sizes. We ask ourselves for any given mouse – do you see a 400 base pair fragment or a 1200 base pair fragment or both? In this example, it is clear that there are some mice that contain just the 400 base pair fragment. That means that the two chromosomes of the PSD-95 gene are the normal kind, they’re not genetically modified. This is DNA from a normal mouse. Now, we can see some other mice that contain a 400 base pair fragment and a 1200 base pair fragment in the same lane. This tells us that there is a normal chromosome, as well as a mutated chromosome. These are heterozygous, just like the parental mouse. We can see there is another lane in there, where there is only the 1200 base pair fragment. This means that there is no normal, wild-type chromosome, and there must only be two copies of the mutant chromosome. This is a homozygous mutation. These mice contain no normal copies of PSD-95, and they contain two copies of the mutated gene. These are what we commonly call “knockout miceâ€, and these are the ones we are most interested in studying.
knock out, knockout, ko, agouti, mouse, model, organism, embryonic, stem, cells, dna, vector, homologous, recombination, blastosyst, neomicin, gangcyclovir, psd95, postsynaptic, density, seth grant,
- ID: 897
- Source: DNALC.G2C
Related Content
1213. PSD-95 and Learning
Professor Seth Grant highlights PSD95 as an important example of a protein associated with a neurotransmitter receptor that affects learning.
1215. PSD-95 Mutations
In studies of PSD95, Professor Seth Grant's group showed that memories are formed when neurotransmitter receptors associate with proteins.
16856. Animation 41: DNA is only the beginning for understanding the human genome.
Mario Capecchi describes proteomics; the large-scale study of protein structure and function. Brian Sauer explains gene knock outs.
1363. Genes for memory (lesson)
Students will experiment with an interactive animation to compare mutant and wild-type mice in a water maze. They will analyze data and discuss findings of a research paper.
16869. Problem 41: DNA is only the beginning for understanding the human genome.
Experiment with gene knock outs.
1712. Mouse (Mus musculus)
Mice are small, easy to keep, and complete a generation in only ten weeks. They are also rather closely related to human beings.
548. Model Center
Model organisms share with humans many key biochemical and physiological functions that have been conserved (maintained) by evolution.
1390. Genes for Learning and Memory
An interactive chromosome map of the genes and loci associated with learning and memory.
15060. Homologous recombination, Mario Capecchi
Mario Capecchi discusses homologous recombination, the technique he developed to introduce a desired mutation into the DNA of living cells.
1368. DLG4 Gene
Discs, large homolog 4 (DLG4) is a gene associated with learning and memory. The human DLG4 protein is 99% identical to the rat and mouse PSD-95 proteins.