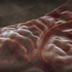
Shotgun sequencing and dealing with repeat sections, 3D animation with basic narration
The private project blew the genome apart into pieces of different sizes and used mathematical models and other maps to reconstruct the genome from these pieces. (DNAi Location: Genome > The Project > Putting it together > Animations > Whole genome shotgun-private)
Shotgun sequencing is the method that was used by the private genome project. Shotgun sequencing requires multiple copies of the genome, which are effectively blown up into millions of small fragments. Each fragment is then sequenced. The small fragments are assembled using an immense amount of computer power to match overlapping sections. The drawback of this method comes when dealing with repeat sequences. Often there is no way of knowing how long the repeat sequence is; or in which of many different possible positions the fragments overlap. Even the incredibly powerful software used to shotgun sequence the human genome couldn't cope with this. So Celera, the private company which relied on this approach, had to use the public data to fill in the gaps left by the repeats.
shotgun sequencing,powerful software,computer power,genome project,human genome,fragments,drawback,narration,fragment,private company,gaps,sequences,animation
- ID: 15537
- Source: DNALC.DNAi
- Download: MPEG 4 Video
Related Content
15477. The public Human Genome Project: mapping the genome, sequencing, and reassembly. 3D animation.
The public Human Genome Project: mapping the genome, sequencing, and reassembly.
15367. Using data from the public project, Craig Venter
Craig Venter, leader of the private effort at Celera Genomics, speaks about his company's reliance on the public data for reassembly of the Celera sequence.
15365. Whole genome shotgun, Craig Venter
Craig Venter, the leader of the private genome effort, talks about the "whole genome shotgun" technique that was used by Celera Genomics to sequence the human genome.
15538. Genome Sequencing: Shotgun technique, 3D animation with no audio
Genome Sequencing: Shotgun technique.
16812. Animation 39: A genome is an entire set of genes.
James Watson describes sequencing the human genome using markers and BACs, and Craig Venter explains using cDNA libraries, ESTs, and shotgun sequencing.
15340. The Human Genome Project was more than just sequencing, Ari Patrinos
Ari Patrinos, director of the US Department of Energy's sequencing effort, talks about the public genome project's aims that extended beyond those of the private project.
15479. Sanger method of DNA sequencing, 3D animation with narration
The DNA sequencing method developed by Fred Sanger forms the basis of automated "cycle" sequencing reactions today.
15169. The public sequencing process, John Sulston
Nobel Laureate John Sulston, a key figure in the UK sequencing effort, talks about breaking DNA apart so that the sequence can be reassembled.
15291. Finding genes in the human genome, Ewan Birney
For the first draft of the genome sequence, both teams were working to identify the number of human genes. Here, Ewan Birney, a "numbers man" from the public genome project, explains how genes can be recognized and the data from the genome project used.
15290. Finding genes, Ewan Birney
Ewan Birney talks about finding genes.