Molecules for Memory
Communication in brain cells is guided by interactions between genes and biochemicals at the synapse. These interactions can lead to the formation of new synapses.
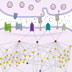
We can study memory from a number of perspectives: we can explore the interactions of a network of genes, examine how biochemicals relay signals, track synapse changes in cells, trace anatomical changes in the brain, or look for changes in behavior over a period of time. Whichever approach we choose, we are ultimately examining the same phenomenon, but at a particular level of organization. In Molecules for Memory, we examine memory at the molecular level. Communication in brain cells is guided by interactions between genes and biochemicals at the synapse. These interactions can lead to the formation of new synapses, which can facilitate long-term learning. Elsewhere on Genes to Cognition Online, you can explore memory formation at the level of cells, brain anatomy, and behavior. The animation begins with the arrival of an action potential at the presynaptic terminal and ends with the formation of a new synapse. The initial focus is with AMPA and NMDA receptors. The formation of a memory in the neuron begins at the synapse. Memories begin to form when a series of action potentials arrive at the presynaptic terminal. The action potential causes a sudden shift in the electrical potential across the membrane. A number of different electrically charged ions rush across the membrane – among these are calcium ions, which activate vesicles in the presynaptic terminal. These vesicles contain the neurotransmitter, glutamate. The vesicles descend to the synaptic cleft and fuse with the membrane. This causes the neurotransmitter to spill out across synaptic cleft. There are a number of glutamate receptors on the postsynaptic neuron, here we can see NMDA and AMPA receptors. They react differently to the glutamate. AMPA receptors respond immediately to glutamate. They open sodium and potassium ion channels in the postsynaptic neuron. When sodium enters the postsynaptic terminal, it depolarises the postsynaptic membrane. NMDA receptors are slower to respond. They respond after the membrane has depolarized. When glutamate binds to the NMDA receptor, the NMDA receptor changes its shape. The change of shape forces a magnesium ion out of the channel, effectively unblocking the channel. Calcium ions, normally found outside the cell, race into the inside of the postsynaptic terminal. Calcium binds to a protein called calmodulin. In turn, calmodulin activates other proteins – here we can see calmodulin activating a protein called CaMKII. CaMKII is a kinase, which performs some very important functions in the cell. Kinases are enzymes, and these enzymes have a very specialized role: it is to add a phosphate group onto other proteins. When a kinase becomes activated, it adds a phosphate onto other proteins, which are called protein substrates. As a result of adding these phosphate groups, there other proteins are now altered. Here we can see CaMKII activating a number of proteins. Many of these activated proteins activate other proteins. If we take a closer look at the synapse, we can follow one of these protein-protein interactions. Here, we see how calcium influx causes a series of reactions that results in activation of the cAMP protein kinase. The activated cAMP protein kinase moves to the nucleus of the postsynaptic cell, where it binds to another protein called CREB. CREB controls transcription of DNA. When activated, CREB transcribes DNA in the cell nucleus to produce RNA. RNA travels back to the synapse, where it synthesizes new proteins to change the structure of the synapse. Changes includes growth of new synapses, which are thought to be the basis of long-term memory. The molecular processes that lead to the generation of these new synapses are not yet known.
molecules for memory, learning, genes, biochemicals, glutamate, CamKII, cAMP, CREB, potassium ion channels, ampa receptors, nmda receptors, calcium ions, postsynaptic neuron, action potentials, synapses,
- ID: 1277
- Source: DNALC.G2C
Related Content
550. The Neural Code
Cognitive information is encoded in patterns of nervous activity and decoded by molecular listening devices at the synapse. Professor Seth Grant explains how different patterns of neural firing are critical to cognition.
1390. Genes for Learning and Memory
An interactive chromosome map of the genes and loci associated with learning and memory.
1365. CREB1 Gene
The cAMP response element-binding protein 1 (CREB1) gene is a CREB activator and has been found to facilitate long-term memory formation.
1997. Learning and memory
Learning and memory are two intimately linked cognitive processes that stem from interactions with the environment (experience).
1366. CREB2 Gene
CAMP response element-binding protein 2(CREB2) is also known as Activating Transcription Factor 2 (ATF2). CREB2 is a CREB repressor, which means it inhibits long-term memory formation.
1212. NMDA Receptors and Learning (1)
Professor Seth Grant explains that NMDA receptors are important to forming memories - if we block NMDA receptors, we can block learning.
1100. Glutamate Receptors
Professor Graham Collingridge describes the glutamate receptor, AMPA, the workhorse receptor for communicating information.
811. The Glutamate System
Professor Trevor Robbins describes some of the key functions of the excitatory glutamate system, which is integral to information processing and long-term potentiation.
1101. AMPA and NMDA Receptors
Professor Graham Collingridge describes the roles played by NMDA and AMPA receptors in long-term potentiation (LTP).
916. GABRB3 Gene
GABA is the main inhibitory neurotransmitter in the adult brain. GABRA3 is a candidate gene for autism.